"Comparisons of molecules (proteins, DNA) of
various species provide independent and compelling support for the
hypothesis of biological macro-evolution"
For the Affirmative, David
E. Thomas
Round 1, Posted August 16th, 2002
Modified September 27th, 2002 (look for * Footnote)
INTRODUCTION
Evolution is descent with modification - for example, the
hypothesized descent of birds from certain theropod dinosaurs. When
sons and daughters are born slightly different from their parents
(micro-evolution), heredity and descent are involved.
"Macro-evolution" means that over time, descent produces new species,
genera, families, orders, classes, phyla, and kingdoms.
Charles Darwin had poor models of descent at best. His theory of
heredity involved small particles, "gemmules," which flowed through
the blood to collect in sexual organs for transmission to the next
generation (Darwin 1868). Although Darwin's genetic models were
incorrect, he was right about one thing -existing
species can evolve into new species.
If this essay were to cover all of the many examples in which
molecular comparisons support evolution, it would be thousands of
pages long. This evidence could have turned out to be
incompatible with common descent in any number of ways. The
biomolecular revolution could easily have demolished evolution,
exposing it as a "fantasy" based on superficial similarities of a few
species. But, again and again, the effects of descent with
modification have been confirmed.
During the last century's explosion of biotechnology, molecular
biologists have observed only one mechanism for similarity in
various organisms' genetic material (DNA) or its expressions
(proteins). That mechanism is descent - heredity. A man may
protest that he's not the father of a given child, but if that
child's DNA has certain similarities to the man's, there is no doubt
he is the father. DNA can establish paternity of individuals, and it
can also indicate paternity of species - macro-evolution.
PROTEIN COMPARISONS
Cytochrome c is a vital protein used at the most basic level of
cellular electron transport. It is one of a number of
ubiquitous proteins - proteins that serve the exact same
function in all species. All surviving organisms require a copy of
cytochrome c, or an equivalent protein. While these proteins vary
from species to species, they all have the same functional shape.
Indeed, yeast cells function just fine with human cytochrome c
engineered into their cells (Tanaka 1988). There are untold
trillions of possible combinations of the 100 or so amino
acids involved that make perfectly functional copies of
cytochrome c. Only one mechanism has been observed to produce genetic
similarity: heredity. Without heredity involved, there is no
reason to expect the proteins of any two organisms to be similar.
Evolution demands similarities in the genes and proteins of
species thought to have recently evolved from a common ancestor.
Since biologists think that humans and chimps both descended from an
ancestor who lived just five to eight million years ago, the
molecules of inheritance (DNA) and the DNA's expressed
proteins should be quite similar between these two species.
Conversely, one would expect many more differences between creatures
related more distantly, such as humans and turtles.
In fact, there is no difference between the cytochrome c's
of human and chimp. Human cytochrome c differs from a rhesus monkey's
by just one amino acid, and from an erythrocebus patas monkey's by a
different one (Dayhoff 1979). But, humans differ from whales at ten
different cytochrome c sites, at 15 for turtles, and so on (Figure
1). There is a "Message" in these proteins: species thought to be
closely related turn out to have proteins that are also
closely related. If human cytochrome sequences were completely
different from those of the apes, or even all other creatures,
evolution would have collapsed overnight. Instead, the molecules were
in perfect accord with evolutionary expectations - independent and
compelling confirmation.

Figure 1 - Comparisons of cytochrome c
sequences.
DNA COMPARISONS
There's an even deeper level of confirmation. It turns out
that there are from one to six codons (DNA nucleotide triplets)
corresponding to each of the twenty amino acids. There are thus
many DNA combinations that can code for the same amino
acid sequence. There is no reason to expect that organisms with
similar protein sequences should have similar DNA
sequences, except for heredity. Figure 2 shows a comparison of
human and mice genes and amino acids for cytochrome c. In the absence
of a mechanism like heredity, the probability that humans and mice
share identical codons for 78 of the 105 amino acids in the protein
is astonishingly small: 7.6*10-34 (NCBI 2002, Hinrichs
2000, Theobald 2002).

Figure 2. DNA Redundancies in Man and Mouse
MULTI-GENE COMPARISONS
Cytochrome c phylogenies are sometimes imperfect, since they are
based on a single gene, and mutations can occasionally reverse
previous mutations. In Figure 1, the lamprey [cytochrome c]
appears closer to humans than does that of tuna fish, contrary to
evolutionary expectations*. Biologists have addressed this problem by
looking at more and more genes. Some genes evolve faster than
others, some slower. Of course, the fact that some genes change
faster or slower than others doesn't disprove evolution.
A recent study (Kumar and Hedges 1998) involved hundreds of
genes, for dozens of species. In Figure 3, the 50 genes used to
compare old world monkeys to humans have a strong "peak" indicating
23 million years (MYa) since the last common ancestor of these
species. The bell-shaped curve centered at 23 MYa is nothing less
than a strong biotic "signal." When 107 genes of primates and rabbits
are compared, these differences correspond to 90 MYa since the
primate/rabbit common ancestor. The "message" is clear: humans are
genetically much closer to old world monkeys than they are to
rabbits. This accords perfectly with evolutionary expectations:
the last common ancestor of primates and rabbits is much farther
back in time than the last common ancestor of humans and old world
monkeys. Had the genes turned out otherwise, Darwin's "fantasy"
would have evaporated.
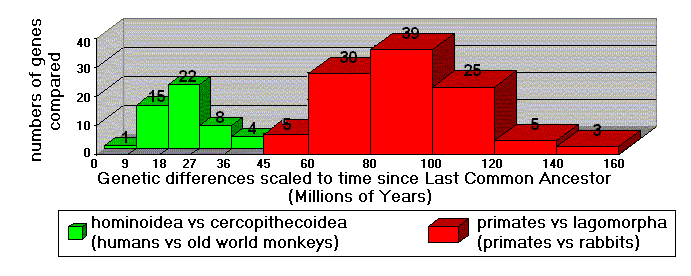
Figure 3. Comparisons of hundreds of genes. (Data from Kumar
& Hedges1998)
GENETIC CODE COMPARISONS
Most organisms have the very same genetic code, the
association of DNA/RNA triplets with specific amino acids. For
example, in most organisms, UGA is a "stop" codon, and specifies the
end of a protein. But in some Mycoplasma bacteria, UGA codes for the
amino acid tryptophan instead (Inamine 1990). Several natural
hypotheses for the evolution of non-standard genetic codes have been
advanced and published (Osawa and Jukes 1995). If all creatures are
descended from one or a few ancestors, then such variant genetic
codes should also display a pattern of derived similarities
and differences, just like those in protein sequence comparisons. And
so they do (Knight 2001, Miller 2001). The real significance of the
few non-standard code exceptions is that they demonstrate that
organisms can indeed exist with their own unique genetic code
translations. There is nothing to stop a Designer from giving any
species such as humans a special genetic code, unlike that of any
other creature (Hofstadter 1982). The 2.3*1069 possible
genetic codes are more than enough for each baramin (kind), even each
species, to have its own special code. This, again, could have
dramatically falsified evolution.
NON-FUNCTIONAL DNA COMPARISONS
Some of the most compelling molecular evidence for
evolution involves inheritance of non-functional DNA, like
introns (Figure 2). While most vertebrates synthesize their own
vitamin C (ascorbic acid), some primates can't. When sequences of
vertebrates are compared, we find that primates do possess genes for
making vitamin C. In humans and apes, however, these genes are
non-functional. A mutation has destroyed the "recipe," but
it's not a "harmful" mutation because primates get vitamin C in their
diets. The mutations blocking human production of this enzyme are
identical to those in the apes. It's unlikely the same
mistakes happened independently, in exactly the same positions; it's
more likely that humans and apes inherited this error from their
common ancestor. (Max 2000). Additionally, curious DNA fragments
called "transposons" ("jumping genes") can inject their DNA randomly
into the genome (Armstrong 2000). Once inserted, these genetic
fragments are then inherited. Some SINE (short interspersed elements)
transposons are routinely used for paternity suits (Novick 1993,
1995); others show the same genetic elements, randomly inserted into
the same locations, in both humans and chimps (Sawada 1985).
NEW SPECIES FROM SIMPLE MUTATIONS
Nylon wasn't invented until 1937. When scientists found a new
species of bacteria feasting on nylon byproducts, they became quite
interested. When they compared old and new strains, they found the
new strain must have nylon byproducts to live, and that its
novel nylon-digesting abilities were due to a frame shift mutation in
one gene. In a frame shift, a mutation of just one nucleotide can
push the whole sequence by a notch, creating a brand-new protein. In
this case, the protein reacted to nylon products, and suddenly gave
the bacteria a whole new niche. The novel proteins we see in real
life do indeed appear to be due to inherited changes in DNA, just as
evolution demands (Ohno 1984, Thomas 2002).
CONCLUSION
In each of these cases, things could have turned out in thousands
of different ways, each with the potential to disprove evolution. But
that's not what happened. Comparisons of molecules provide
independent and compelling support for macro-evolution.
*Two words were added by agreement of the debators to clarify
both a mistake in Mr. Thomas' first submission and Mr. ReMine's
response to that sentence.
REFERENCES
W.P. Armstrong, (2000). "Transposons (Jumping Genes)," http://waynesword.palomar.edu/transpos.htm.
C. R. Darwin (1868). The Variation of Animals and Plants under
Domestication , ISBN: 0801858674.
Margaret Dayhoff (1979) Atlas of Protein Sequence and
Structure, ISBN: 0912466073; (April 1979 edition)
Steve Hinrichs (June 2000). "Compelling Data for Common Descent
from Matching Redundant DNA Sequences," http://members.aol.com/SHinrichs9/descent/descent.htm.
Douglas Hofstadter (1982). "Is the genetic code an arbitrary one,
or would another code work just as well?", Scientific American, March
1982, 18-29.
J.M. Inamine et. al. (1990). "Evidence that UGA is read as
tryptophan rather than stop by Mycoplasma ...," Journal of
Bacteriology 172;504-506.
Robin Knight et. al., (2001) "Rewiring The Keyboard: Evolvability
Of The Genetic Code," Nature Reviews - Genetics. 2: 49-58.
S. Kumar and S. Hedges (1998). "A molecular timescale for
vertebrate evolution," Nature, Vol. 392, 30 April 1998, pp.
917-920.
Edward Max (2000). "Plagiarized Errors and Molecular Genetics,"
<http://www.talkorigins.org/faqs/molgen/>.
Kenneth R. Miller (2001), http://www.ncseweb.org/resources/articles/6945_km-3.pdf.
NCBI (2002). "Somatic cytochrome c Gene," Accession Number
M22877
(Human), X01756
(mouse).
G. E. Novick, et. al., (1993). "The use of polymorphic Alu
insertions in human DNA fingerprinting," EXS. 67: 283-291.
[PubMed]
G. E. Novick, et. al., (1995) "Polymorphic human specific Alu
insertions ...," Electrophoresis 16: 1596-1601. [PubMed]
Susumu Ohno (1984) "Birth of a unique enzyme ...," Proc. Natl.
Acad. Sci. USA, Vol. 81, pp. 2421-2425, April 1984.
S. Osawa, T. H. Jukes (1995) "On Codon Reassignment," Journal of
Molecular Evolution, 41:247-249.
I. Sawada, et al. (1985) "Evolution of Alu family repeats since
the divergence of human and chimpanzee," Journal of Molecular
Evolution 22(316). [PubMed]
Y. Tanaka, et. al. (1988) "Construction of a human cytochrome c
gene and its functional expression in Saccharomyces cerevisiae," J
Biochem (Tokyo) (1988) 103: 954-61.[PubMed]
Douglas Theobald (2002). "The Molecular Sequence Evidence,"
http://www.talkorigins.org/faqs/comdesc/section4.html.
David E. Thomas (July 2002) "Evolution and Information: The Nylon
Bug," http://www.nmsr.org/nylon.htm.

This document was last modified on 09/26/2003
http://www.nmsr.org/round1a.html
Hits from Inception thru August 1st, 2003: 1136
NMSR Debate
Home 
TCCSA Debate
Home 
NMSR
Home



